Institute for Biomedicine - News & Events - Monitoring of SARS-CoV-2 mutations
Monitoring of SARS-CoV-2 mutations
Eurac Research supports the Health Authority in the sequencing of the virus genome.
- Deutsch
- English
- Italiano
In mid-January 2021, national and local health authorities were faced with a new phase of the pandemic: the increasing spread of new mutations of the virus.
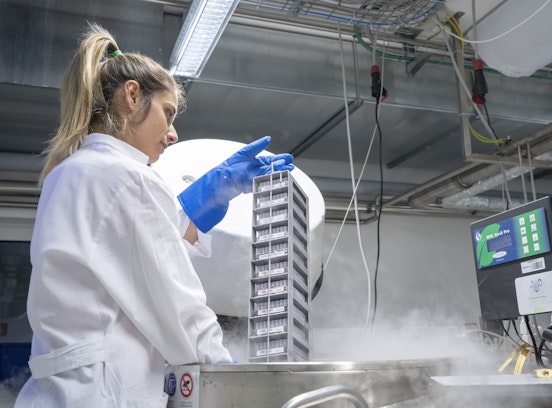
The Istituto Superiore di Sanità is coordinating the regions to analyse the genome of SARS-CoV-2 in some of the swabs carried out on Italian territory, in order to trace and identify mutant strains. This type of monitoring on SARS-CoV-2 involves the sequencing of the entire viral genome, a different process from the molecular analysis usually performed on swabs. Normally, a swab is used to search for the presence of the virus by identifying and quantifying specific fragments of SARS-CoV-2 genetic material. Sequencing, on the other hand, involves reading the entire viral genome to identify possible differences and variants. This technique employs more complex time and technology than molecular swab analysis. The microbiology and virology laboratory of the South Tyrolean Health Authority was also contacted in order to meet the need of the Istituto Superiore di Sanità to analyse the variants. The Eurac Research laboratories are equipped for this type of analysis because they have been dealing with human genome sequencing for years. They also have at their disposal the Next Generation Sequencing (NGS) technology, which was made available to the South Tyrolean Health Authority in order to quickly start sequencing the first samples in which SARS-CoV-2 was found. The design of this particular initiative - which deviates from standard activities - required experimental planning and careful preparation of materials coordinated by the scientific staff of the Institute of Biomedicine of Eurac Research. Once the preparation phase has been completed, the analysis of a first slot of samples selected by the Health Authority will start in the second week of March. Initially, the genetic material of about 50 samples per month will be analysed; after an initial set-up phase, the analysis capacity can be optimised to process up to 200 samples per month. This new form of collaboration is part of a year in which the management of the health emergency has required the sudden acquisition of many materials and instruments to face different needs from the ordinary health activity.
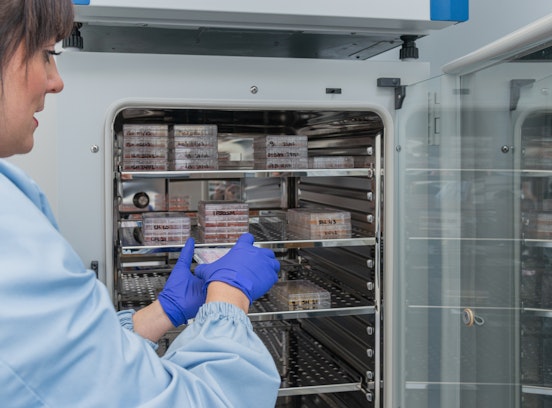
Since the beginning of the pandemic, Eurac Research has collaborated with the Health Authority offering logistical and scientific support, providing equipment and specialized personnel. In the first weeks of the emergency, in March 2020, laboratory equipment and instruments, as well as laboratory staff, were made available to the Bolzano Hospital. In order to facilitate the analysis of the numerous molecular swabs, the Institute of Biomedicine also temporarily allocated an automatic extractor to the Health Authority's Microbiology and Virology Laboratory to quickly process 96 samples at a time. In addition, at the beginning of 2021, three freezers cooling down to -80 degrees were provided for the storage of the new RNA vaccines, facilitating the storage of the first vaccine supplies.